
Chinese Journal of Applied Chemistry ›› 2022, Vol. 39 ›› Issue (1): 3-17.DOI: 10.19894/j.issn.1000-0518.210479
• Review • Previous Articles Next Articles
Protein Sequence Design Using Generative Models
WU Qing-Lin1, REN Yu-Bin2, ZHAI Xiao-Wei1, CHEN Dong1(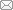 ), LIU Kai2(
), LIU Kai2( )
)
- 1.College of Energy Engineering,Zhejiang University,Hangzhou 310012,China
2.Department of Chemistry,Tsinghua University,Beijing 100084,China
-
Received:2021-09-26Accepted:2021-11-11Published:2022-01-01Online:2022-01-10 -
Contact:Dong CHEN,Kai LIU -
About author:kailiu@mail.tsinghua.edu.cn
kailiu@mail.tsinghua.edu.cn
-
Supported by:the National Natural Science Foundation of China(21878258);Zhejiang Provincial Natural Science Foundation of China(Y20B060027)
CLC Number:
Cite this article
WU Qing-Lin, REN Yu-Bin, ZHAI Xiao-Wei, CHEN Dong, LIU Kai. Protein Sequence Design Using Generative Models[J]. Chinese Journal of Applied Chemistry, 2022, 39(1): 3-17.
share this article
Add to citation manager EndNote|Ris|BibTeX
URL: http://yyhx.ciac.jl.cn/EN/10.19894/j.issn.1000-0518.210479

Fig. 2 Design sequence for a given backboneA. General steps[38]; B. Two evaluation methods based on energy minimization or probability maximization[36]
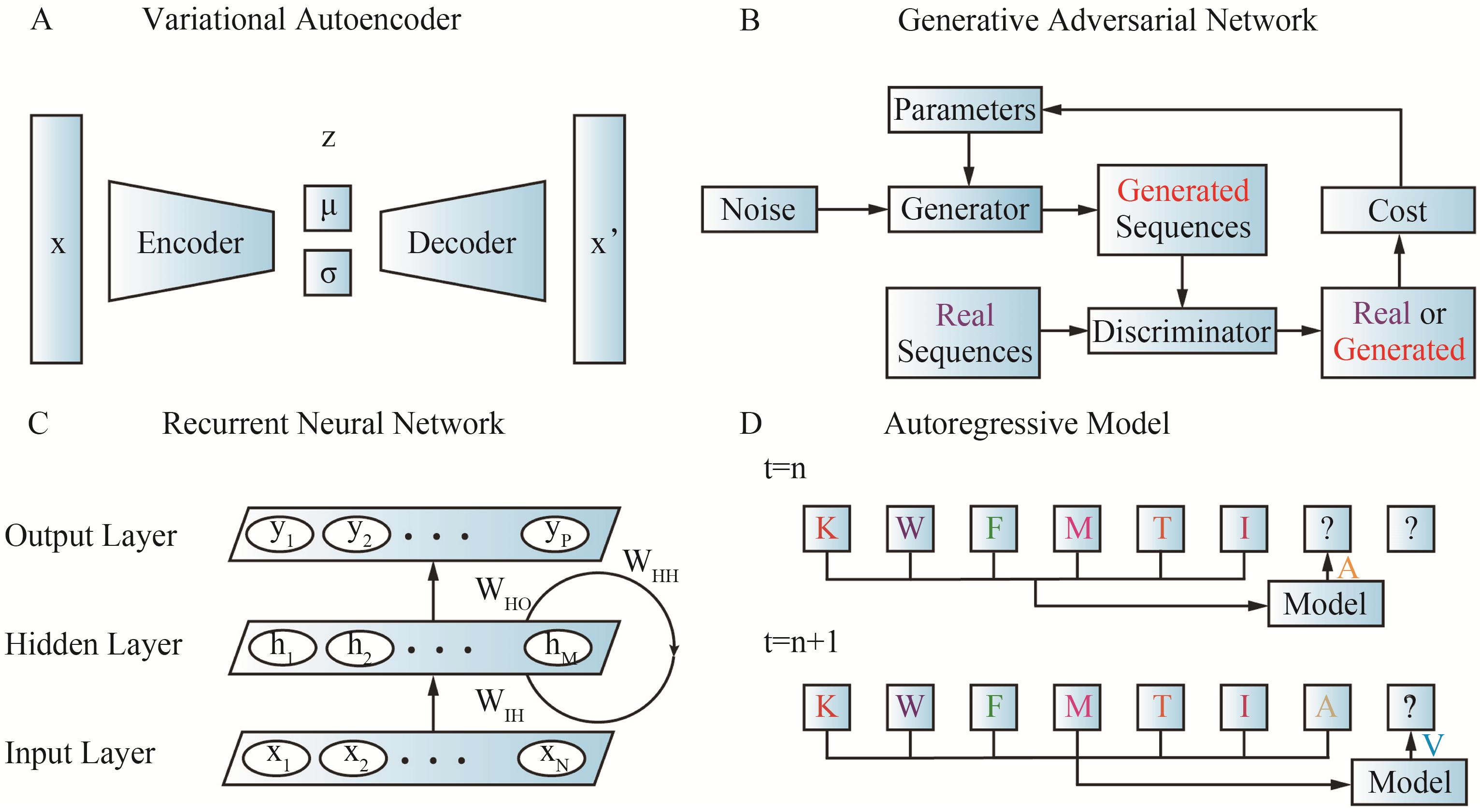
Fig.3 Four commonly used generative models in protein sequence design(A) Variational autoencoder (VAE)[24]; (B) Generative adversarial network (GAN)[24]; (C) Recurrent neural network (RNN)[48]; (D) Autoregressive model (ARM)[24]
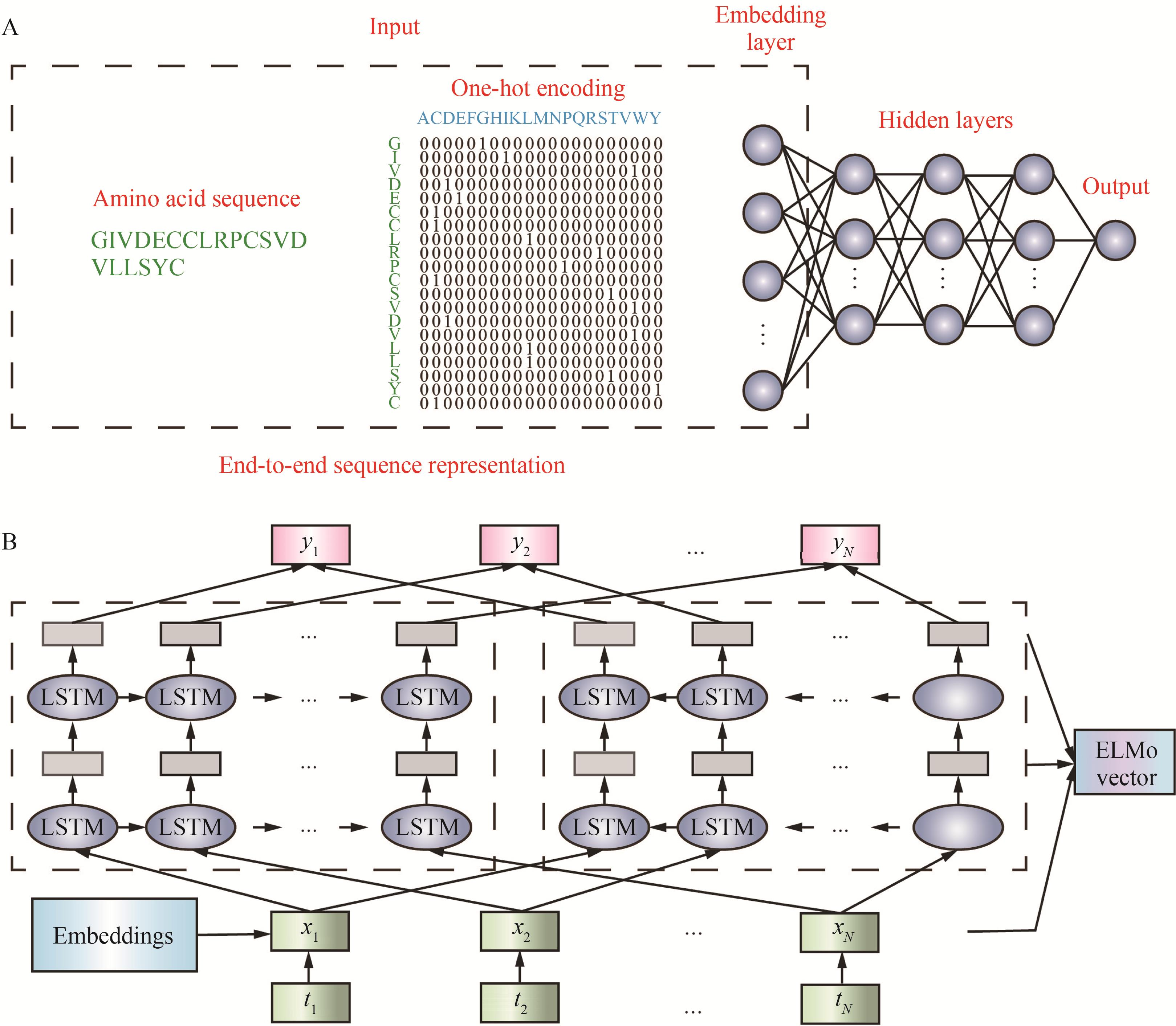
Fig. 4 Two commonly used methods for protein sequence representation(A) End-to-end sequence representation[71]; (B) Sequence representation by ELMo model[71]
表示方法 Representation methods | 关键机制 Key mechanisms | 优点 Advantages | 缺点 Disadvantages | 应用 Applications |
|---|---|---|---|---|
端到端学习 End-to-end learning | 独热编码 One-hot encoding | 输入直达输出; 获得氨基酸相似性和差异性 Directly from input to output; capture the physicochemical similarities and differences between amino acids | 需要大量高质量有标签的数据 Need a lot of high quality labeled datas | 蛋白质功能预测、分类等 Protein function prediction, classification, etc. |
迁移学习 Transfer Learning | LSTM(ELMo、BERT) | 利用无标签数据;效果好 Using unlabeled data; good performance | 二级结构、蛋白质功能、 蛋白质演化、相互作用等 Secondary structure, protein function, protein evolution, interaction, etc. | |
上下文无关学习 Non-contextual embedding learning | word2vec等 word2vec et al | 性能不佳 Poor performance | 蛋白质分类、功能识别等 Protein classification, functional recognition, etc. | |
其它学习 Other representation methods | GCN、基于特征提取的方法 GCN, feature extraction based method | 应用局限在特定领域 Applications are limited to specific domains | 蛋白质组、蛋白药物、 蛋白质结构等特定领域 Roteome, protein medicine, protein structure and other specific fields |
Table1 Comparison of four types of protein sequence representation
表示方法 Representation methods | 关键机制 Key mechanisms | 优点 Advantages | 缺点 Disadvantages | 应用 Applications |
|---|---|---|---|---|
端到端学习 End-to-end learning | 独热编码 One-hot encoding | 输入直达输出; 获得氨基酸相似性和差异性 Directly from input to output; capture the physicochemical similarities and differences between amino acids | 需要大量高质量有标签的数据 Need a lot of high quality labeled datas | 蛋白质功能预测、分类等 Protein function prediction, classification, etc. |
迁移学习 Transfer Learning | LSTM(ELMo、BERT) | 利用无标签数据;效果好 Using unlabeled data; good performance | 二级结构、蛋白质功能、 蛋白质演化、相互作用等 Secondary structure, protein function, protein evolution, interaction, etc. | |
上下文无关学习 Non-contextual embedding learning | word2vec等 word2vec et al | 性能不佳 Poor performance | 蛋白质分类、功能识别等 Protein classification, functional recognition, etc. | |
其它学习 Other representation methods | GCN、基于特征提取的方法 GCN, feature extraction based method | 应用局限在特定领域 Applications are limited to specific domains | 蛋白质组、蛋白药物、 蛋白质结构等特定领域 Roteome, protein medicine, protein structure and other specific fields |

Fig.6 Two methods for optimizing protein sequences(A) Feedback generative adversarial network (FBGAN)[83]; (B) Agent-environment interactions in reinforcement learning (RL)[88]
| 1 | HUANG P S, BOYKEN S E, BAKER D. The coming of age of de novo protein design[J]. Nature, 2016, 537(7620): 320-327. |
| 2 | GAINZA-CIRAUQUI P, CORREIA B E. Computational protein design—the next generation tool to expand synthetic biology applications[J]. Curr Opin Biotechnol, 2018, 52: 145-152. |
| 3 | CHEVALIER A, SILVA D A, ROCKLIN G J, et al. Massively parallel de novo protein design for targeted therapeutics[J]. Nature, 2017, 550(7674): 74-79. |
| 4 | KAMERZELL T J, MIDDAUGH C R. Prediction machines: applied machine learning for therapeutic protein design and development[J]. J Pharm Sci, 2021, 110(2): 665-681. |
| 5 | SILVA D A, YU S, ULGE U Y, et al. De novo design of potent and selective mimics of IL-2 and IL-15[J]. Nature, 2019, 565(7738): 186-191. |
| 6 | LI J, LI B, SUN J, et al. Engineered near-infrared fluorescent protein assemblies for robust bioimaging and therapeutic applications[J]. Adv Mater, 2020, 32(17): 2000964. |
| 7 | SUN J, LI B, WANG F, et al. Proteinaceous fibers with outstanding mechanical properties manipulated by supramolecular interactions[J]. CCS Chem, 2021, 3(6): 1669-1677. |
| 8 | XIAO L, WANG Z, SUN Y, et al. An artificial phase-transitional underwater bioglue with robust and switchable adhesion performance[J]. Angew Chem Int Ed, 2021, 60(21): 12082-12089. |
| 9 | ANAND-ACHIM N, EGUCHI R R, MATHEWS I I, et al. Protein sequence design with a learned potential[J]. bioRxiv, 2021,DOI: 10.1101/2020.01.06.895466 |
| 10 | LI Y, LI J, SUN J, et al. Bioinspired and mechanically strong fibers based on engineered non-spider chimeric proteins[J]. Angew Chem Int Ed, 2020, 132(21): 8225-8229. |
| 11 | KUHLMAN B, BRADLEY P. Advances in protein structure prediction and design[J]. Nat Rev Mol Cell Biol, 2019, 20(11): 681-697. |
| 12 | CHEN I M A, MARKOWITZ V M, CHU K, et al. IMG/M: integrated genome and metagenome comparative data analysis system[J]. Nucleic Acids Res, 2017, 45(D1): D507-D516. |
| 13 | WESTBROOK J D, BURLEY S K. How structural biologists and the protein data bank contributed to recent FDA new drug approvals[J]. Structure, 2019, 27(2): 211-217. |
| 14 | LECUN Y, BENGIO Y, HINTON G. Deep learning[J]. Nature, 2015, 521(7553): 436-444. |
| 15 | TUNYASUVUNAKOOL K, ADLER J, WU Z, et al. Highly accurate protein structure prediction for the human proteome[J]. Nature, 2021, 596(7873): 590-596. |
| 16 | YANG J Y, ANISHCHENKO I, PARK H, et al. Improved protein structure prediction using predicted interresidue orientations[J]. Proc Natl Acad Sci USA, 2020, 117(3): 1496-1503. |
| 17 | STROKACH A, BECERRA D, CORBI-VERGE C, et al. Fast and flexible protein design using deep graph neural networks[J]. Cell Syst, 2020, 11(4): 402-4114. |
| 18 | WEI K Y, MOSCHIDI D, BICK M J, et al. Computational design of closely related proteins that adopt two well-defined but structurally divergent folds[J]. Proc Natl Acad Sci USA, 2020, 117(13): 7208-7215. |
| 19 | BAEK M, DIMAIO F, ANISHCHENKO I, et al. Accurate prediction of protein structures and interactions using a three-track neural network[J]. Science, 2021, 373(6557): 871-876. |
| 20 | CUNNINGHAM J M, KOYTIGER G, SORGER P K, et al. Biophysical prediction of protein-peptide interactions and signaling networks using machine learning[J]. Nat Methods, 2020, 17(2): 175-183. |
| 21 | HOPF T A, INGRAHAM J B, POELWIJK F J, et al. Mutation effects predicted from sequence co-variation[J]. Nat Biotechnol, 2017, 35(2): 128-135. |
| 22 | RIESSELMAN A J, INGRAHAM J B, MARKS D S. Deep generative models of genetic variation capture the effects of mutations[J]. Nat Methods, 2018, 15(10): 816-822. |
| 23 | WANG D D, LE O Y, XIE H R, et al. Predicting the impacts of mutations on protein-ligand binding affinity based on molecular dynamics simulations and machine learning methods[J]. Comput Struct Biotechnol J, 2020, 18: 439-454. |
| 24 | WU Z, JOHNSTON K E, ARNOLD F H, et al. Protein sequence design with deep generative models[J]. Curr Opin Chem Biol, 2021, 65: 18-27. |
| 25 | SINAI S, KELSIC E, CHURCH G M, et al. Variational auto⁃encoding of protein sequences[J]. arXiv e⁃prints, 2017: arXiv:. |
| 26 | Bitard⁃Feildel T. Navigating the amino acid sequence space between functional proteins using a deep learning framework[J]. bioRxiv, 2021: 2020.11. 09.375311. DOI: https://doi.org/10.1101/2020.11.09.375311 |
| 27 | REPECKA D, JAUNISKIS V, KARPUS L, et al. Expanding functional protein sequence spaces using generative adversarial networks[J]. Nat Mach Intell, 2021, 3(4): 324-333. |
| 28 | MÜLLER A T, HISS J A, SCHNEIDER G. Recurrent neural network model for constructive peptide design[J]. J Chem Inf Model, 2018, 58(2): 472-479. |
| 29 | SHIN J E, RIESSELMAN A J, KOLLASCH A W, et al. Protein design and variant prediction using autoregressive generative models[J]. Nat Commun, 2021, 12(1): 1-11. |
| 30 | COLUZZA I. Computational protein design: a review[J]. J Phys: Condens Matter, 2017, 29(14): 143001. |
| 31 | TINBERG C E, KHARE S D, DOU J Y, et al. Computational design of ligand-binding proteins with high affinity and selectivity[J]. Nature, 2013, 501(7466): 212-216. |
| 32 | KOEPNICK B, FLATTEN J, HUSAIN T, et al. De novo protein design by citizen scientists[J]. Nature, 2019, 570(7761): 390-394. |
| 33 | YANG C, SESTERHENN F, BONET J, et al. Bottom-up de novo design of functional proteins with complex structural features[J]. Nat Chem Biol, 2021, 17(4): 492-500. |
| 34 | DAWSON W M, RHYS G G, WOOLFSON D N. Towards functional de novo designed proteins[J]. Curr Opin Chem Biol, 2019, 52: 102-111. |
| 35 | PAN X, KORTEMME T. Recent advances in de novo protein design: principles, methods, and applications[J]. J Biol Chem, 2021, 296: 100558. |
| 36 | NORN C, WICKY B I M, JUERGENS D, et al. Protein sequence design by conformational landscape optimization[J]. Proc Natl Acad Sci USA, 2021, 118(11): e2017228118. |
| 37 | SANDHYA S, MUDGAL R, KUMAR G, et al. Protein sequence design and its applications[J]. Curr Opin Struct Biol, 2016, 37: 71-80. |
| 38 | LIU H Y, CHEN Q. Computational protein design for given backbone: recent progresses in general method-related aspects[J]. Curr Opin Struct Biol, 2016, 39: 89-95. |
| 39 | MAGUIRE J B, HADDOX H K, STRICKLAND D, et al. Perturbing the energy landscape for improved packing during computational protein design[J]. Proteins Struct Funct Bioinf, 2021, 89(4): 436-449. |
| 40 | XIONG P, HU X H, HUANG B, et al. Increasing the efficiency and accuracy of the ABACUS protein sequence design method[J]. Bioinformatics, 2020, 36(1): 136-144. |
| 41 | KINGMA D P, WELLING M. Auto⁃encoding variational bayes[J]. arXiv e⁃prints, 2013: arXiv:. |
| 42 | HAWKINS-HOOKER A, DEPARDIEU F, BAUR S, et al. Generating functional protein variants with variational autoencoders[J]. PLoS Comput Biol, 2021, 17(2): e1008736. |
| 43 | GOODFELLOW I, POUGET-ABADIE J, MIRZA M, et al. Generative adversarial nets[J]. Proc Adv Neural Inf Process Syst, 2014, 27. |
| 44 | MIRZA M, OSINDERO S. Conditional generative adversarial nets[J]. arXiv preprint arXiv, 2014, 1411.1784. |
| 45 | ARJOVSKY M, BOTTOU L. Towards principled methods for training generative adversarial networks[J]. arXiv e⁃prints, 2017: arXiv:. |
| 46 | ARJOVSKY M, CHINTALA S, BOTTOU L. Wasserstein generative adversarial networks[C]//International conference on machine learning. PMLR, 2017: 214-223. |
| 47 | YU L, ZHANG W, WANG J, et al. Seqgan: sequence generative adversarial nets with policy gradient[C]//Proceedings of the AAAI conference on artificial intelligence. 2017. |
| 48 | SALEHINEJAD H, SANKAR S, BARFETT J, et al. Recent advances in recurrent neural networks[J]. arXiv e⁃prints, 2017: arXiv:. |
| 49 | TRAN K, BISAZZA A, MONZ C. Recurrent memory networks for language modeling[J]. arXiv e⁃prints, 2016: arXiv:. |
| 50 | MIROWSKI P, VLACHOS A. Dependency recurrent neural language models for sentence completion[J]. arXiv e⁃prints, 2015: arXiv:. |
| 51 | LE Q V, JAITLY N, HINTON G E. A simple way to initialize recurrent networks of rectified linear units[J]. arXiv e⁃prints, 2015: arXiv:. |
| 52 | SHERSTINSKY A. Fundamentals of recurrent neural network (RNN) and long short-term memory (LSTM) network[J]. Phys D, 2020, 404: 132306. |
| 53 | JÓZEFOWICZ R, ZAREMBA W, SUTSKEVER I. An empirical exploration of recurrent network architectures[C]//International conference on machine learning. PMLR, 2015: 2342-2350. |
| 54 | GRISONI F, SCHNEIDER G. De novo molecular design with generative long short-term memory[J]. Chimia, 2019, 73(12): 1006-1011. |
| 55 | SCHNEIDER P, WALTERS W P, PLOWRIGHT A T, et al. Rethinking drug design in the artificial intelligence era[J]. Nat Rev Drug Discovery, 2020, 19(5): 353-364. |
| 56 | XU M Y, RAN T, CHEN H M. De novo molecule design through the molecular generative model conditioned by 3D information of protein binding sites[J]. J Chem Inf Model, 2021, 61(7): 3240-3254. |
| 57 | CHEN X, MISHRA N, ROHANINEJAD M, et al. Pixelsnail: an improved autoregressive generative model[C]//International Conference on Machine Learning. PMLR, 2018: 864-872. |
| 58 | RIESSELMAN A, SHIN J⁃E, KOLLASCH A, et al. Accelerating protein design using autoregressive generative models[J]. bioRxiv, 2021: 757252. DOI: https://doi.org/10.1101/757252. |
| 59 | BENGIO Y, COURVILLE A, VINCENT P. Representation learning: a review and new perspectives[J]. IEEE Trans Pattern Anal Mach Intell, 2013, 35(8): 1798-1828. |
| 60 | JURTZ V I, JOHANSEN A R, NIELSEN M, et al. An introduction to deep learning on biological sequence data: examples and solutions[J]. Bioinformatics, 2017, 33(22): 3685-3690. |
| 61 | LIU X R, HONG Z Y, LIU J, et al. Computational methods for identifying the critical nodes in biological networks[J]. Briefings Bioinf, 2020, 21(2): 486-497. |
| 62 | HEFFERNAN R, YANG Y D, PALIWAL K, et al. Capturing non-local interactions by long short-term memory bidirectional recurrent neural networks for improving prediction of protein secondary structure, backbone angles, contact numbers and solvent accessibility[J]. Bioinformatics, 2017, 33(18): 2842-2849. |
| 63 | LIU Z, JIN J, CUI Y, et al. DeepSeqPanII: an interpretable recurrent neural network model with attention mechanism for peptide-HLA class II binding prediction[J]. IEEE/ACM Trans Comput Biol Bioinf 2021. DOI:10.1109/TCBB.2021.3074927 |
| 64 | ALLEY E C, KHIMULYA G, BISWAS S, et al. Unified rational protein engineering with sequence-based deep representation learning[J]. Nat Methods, 2019, 16(12): 1315-1322. |
| 65 | ELNAGGAR A, HEINZINGER M, DALLAGO C, et al. ProtTrans: towards cracking the language of life's code through self⁃supervised deep learning and high performance computing[J]. arXiv e⁃prints, 2020: arXiv:. |
| 66 | RIVES A, MEIER J, SERCU T, et al. Biological structure and function emerge from scaling unsupervised learning to 250 million protein sequences[J]. Proc Natl Acad Sci USA, 2021, 118(15): e2016239118. |
| 67 | RAO R, LIU J, VERKUIL R, et al. Msa transformer[J]. bioRxiv, 2021: 2021.02.12.430858. DOI: https://doi.org/10.1101/2021.02.12.430858. |
| 68 | SERCU T, VERKUIL R, MEIER J, et al. Neural Potts Model[J]. bioRxiv, 2021: 2021.04.08.439084. DOI: https://doi.org/10.1101/2021.04.08.439084. |
| 69 | ELABD H, BROMBERG Y, HOARFROST A, et al. Amino acid encoding for deep learning applications[J]. BMC Bioinf, 2020, 21(1): 1-14. |
| 70 | PETERS M E, NEUMANN M, IYYER M, et al. Deep contextualized word representations[J]. arXiv e⁃prints, 2020: arXiv:. |
| 71 | CUI F F, ZHANG Z L, ZOU Q. Sequence representation approaches for sequence-based protein prediction tasks that use deep learning[J]. Briefings Funct Genomics, 2021, 20(1): 61-73. |
| 72 | HEINZINGER M, ELNAGGAR A, WANG Y, et al. Modeling aspects of the language of life through transfer-learning protein sequences[J]. BMC Bioinf, 2019, 20(1): 723. |
| 73 | CHAROENKWAN P, NANTASENAMAT C, HASAN M M, et al. BERT4Bitter: a bidirectional encoder representations from transformers (BERT)-based model for improving the prediction of bitter peptides[J]. Bioinformatics, 2021, 37(17): 2556-2562. |
| 74 | GREENER J G, MOFFAT L, JONES D T. Design of metalloproteins and novel protein folds using variational autoencoders[J]. Sci Rep, 2018, 8(1): 16189. |
| 75 | VASWANI A, SHAZEER N, PARMAR N, et al. Attention is all you need[C]//Advances in neural information processing systems. 2017: 5998-6008. |
| 76 | SURANA S, ARORA P, SINGH D, et al. PandoraGAN: Generating antiviral peptides using Generative Adversarial Network[J]. bioRxiv, 2021: 2021.02.15.431193. DOI: https://doi.org/10.1101/2021.02.15.431193 |
| 77 | YANG R, WU F, ZHANG C, et al. iEnhancer-GAN: a deep learning framework in combination with word embedding and sequence generativeadversarial net to identify enhancers and their strength[J]. Int J Mol Sci, 2021, 22(7): 3589. |
| 78 | CAPECCHI A, CAI X G, PERSONNE H, et al. Machine learning designs non-hemolytic antimicrobial peptides[J]. Chem Sci, 2021, 12(26): 9221-9232. |
| 79 | MAKHZANI A, SHLENS J, JAITLY N, et al. Adversarial autoencoders[J]. arXiv e⁃prints, 2015: arXiv:. |
| 80 | RUSS W P, FIGLIUZZI M, STOCKER C, et al. An evolution-based model for designing chorismate mutase enzymes[J]. Science, 2020, 369(6502): 440-445. |
| 81 | BROOKES D, PARK H, LISTGARTEN J. Conditioning by adaptive sampling for robust design[C]//International conference on machine learning. PMLR, 2019: 773-782. |
| 82 | ANGERMUELLER C, DOHAN D, BELANGER D, et al. Model-based reinforcement learning for biological sequence design[C]//International conference on learning representations. 2019. |
| 83 | GUPTA A, ZOU J. Feedback GAN for DNA optimizes protein functions[J]. Nat Mach Intell, 2019, 1(2): 105-111. |
| 84 | AMIMEUR T, SHAVER J M, KETCHEM R R, et al. Designing feature-controlled humanoid antibody discovery libraries using generative adversarial networks[J]. bioRxiv, 2020. |
| 85 | WAN C, JONES D T. Protein function prediction is improved by creating synthetic feature samples with generative adversarial networks[J]. Nat Mach Intell, 2020, 2(9): 540-550. |
| 86 | LINDER J, BOGARD N, ROSENBERG A B, et al. A generative neural network for maximizing fitness and diversity of synthetic DNA and protein sequences[J]. Cell Syst, 2020, 11(1): 49-62.e16. |
| 87 | RYPEŚĆ G, LEPAK Ł, WAWRZYŃSKI P. Reinforcement learning for on-line sequence transformation[J]. arXiv preprint arXiv, 2105.14097, 2021. |
| 88 | FRANÇOIS-LAVET V, HENDERSON P, ISLAM R, et al. An introduction to deep reinforcement learning[J]. Found Trends Mach Learn, 2018, 11(3-4): 219-354. |
| 89 | POPOVA M, ISAYEV O, TROPSHA A. Deep reinforcement learning for de novo drug design[J]. Sci Adv, 2018, 4(7): eaap7885. |
| 90 | ZHOU Z P, KEARNES S, LI L, et al. Optimization of molecules via deep reinforcement learning[J]. Sci Rep, 2019, 9(1): 10752. |
| 91 | JEON W, KIM D. Autonomous molecule generation using reinforcement learning and docking to develop potential novel inhibitors[J]. Sci Rep, 2020, 10(1): 1-11. |
| 92 | YU H, SCHRECK J S, MA W. Deep reinforcement learning for protein folding in the hydrophobic⁃polar model with pull moves[C]//Third Workshop on Machine Learning and the Physical Sciences, Vancouver, Canada, 2020. |
| [1] | ZHAO Chang-Li, QIN Ming-Gao, DOU Xiao-Qiu, FENG Chuan-Liang. High Mechanical Stability and Osteogenesis of Chiral Supramolecular Hydrogel Induced by Inorganic Nanoparticles [J]. Chinese Journal of Applied Chemistry, 2022, 39(1): 177-187. |
| [2] | HUAI Meng-Jiao, LIU Tao-Xue-Ting, JIANG Zhong-Yi. Research Progress of Artificial Water Channels Inspired by Aquaporin [J]. Chinese Journal of Applied Chemistry, 2022, 39(1): 99-109. |
| [3] | JAING Fuxiang1, LIU Qiaozhen1, WANG Guo1, CHEN Heru1,2*. Synthesis of Piperidine Derivatives from 1,5-Diols [J]. Chinese Journal of Applied Chemistry, 2013, 30(07): 769-775. |
| [4] | LI Huijing, WANG Haishui*. Effect of Boric Acid on the Crystallization of L-Lysine Monohydrochloride Dihydrate on Self-Assembled Monolayers of L-Cysteine [J]. Chinese Journal of Applied Chemistry, 2012, 29(09): 1041-1045. |
| [5] | LIU Qian, LU Jun-Rui*, XIN Chun-Wei, BAO Xiu-Rong, LIU Yu-Qing, ZHU Shan-Shan, ZOU Min. Synthesis, Characterization and Antibacterial Properties of 5-Chloro -o-hydroxybenzyl Amino Acid Esters [J]. Chinese Journal of Applied Chemistry, 2010, 27(09): 1012-1016. |
| [6] | CAI Dong-Qing1, WU Zheng-Yan1*, WU Lin1, WU Yue-Jin1, CHU Paul K2, YU Zeng-Liang1. Removal of Impurities in Carbon Nanotubes by Low Energy Ion Beam Irradiation [J]. Chinese Journal of Applied Chemistry, 2010, 27(08): 987-989. |
| [7] | FENG Yi-Si1, DONG Wen-Jie1, ZHANG Bo1, SHANG Lin1, TAN Wen-Fei2, XU Hua-Jian1*. Synthesis of the Segment of Natural Cyclic Depsipeptide Stereocalpin A [J]. Chinese Journal of Applied Chemistry, 2010, 27(02): 240-242. |
| [8] | RUAN Xiang-Yuan1*, ZENG Shao-Han2, CAI Meng-Zhao2, XU Jing-Wei3. Surface Morphology of Glutathione Complex with Cadmium and Copper via Atomic Force Microscope [J]. Chinese Journal of Applied Chemistry, 2009, 26(12): 1498-1500. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||

