
应用化学 ›› 2022, Vol. 39 ›› Issue (10): 1475-1487.DOI: 10.19894/j.issn.1000-0518.220032
• 综合评述 • 下一篇
基于框架核酸的细胞表面受体配体相互作用研究进展
郭琳洁1,2,3, 彭红珍2, 李江1,2,3, 王丽华1,2,3, 诸颖1,2,3(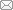 )
)
- 1.中国科学院上海应用物理研究所,中国科学院微观界面与探测重点实验室,上海 201800
2.中国科学院上海高等研究院,基础交叉研究中心,上海 201210
3.中国科学院大学,北京 100049
-
收稿日期:2022-02-11接受日期:2022-04-14出版日期:2022-10-01发布日期:2022-10-05 -
通讯作者:诸颖 -
基金资助:国家自然科学基金(82050005);中国科学院青年创新促进会人才专项(2016236);上海市科委项目(19142202400)
Advances in Receptor‑ligand Interactions on Cell Surface Based on Framework Nucleic Acids
Lin-Jie GUO1,2,3, Hong-Zhen PENG2, Jiang LI1,2,3, Li-Hua WANG1,2,3, Ying ZHU1,2,3( )
)
- 1.Division of Physical Biology,CAS Key Laboratory of Interfacial Physics and Technology,Shanghai Institute of Applied Physics,Chinese Academy of Sciences,Shanghai 201800,China
2.The Interdisciplinary Research Center,Shanghai Advanced Research Institute,Chinese Academy of Sciences,Shanghai 201210,China
3.University of Chinese Academy of Sciences,Beijing 100049,China
-
Received:2022-02-11Accepted:2022-04-14Published:2022-10-01Online:2022-10-05 -
Contact:Ying ZHU -
About author:zhuying@zjlab.org.cn
-
Supported by:the National Natural Science Foundation of China(82050005);the Youth Innovation Promotion Association of CAS(2016236);the Shanghai Municipal Science and Technology Commission(19142202400)
摘要:
细胞表面受体与配体之间的特异性相互作用在细胞生物学过程中起着重要作用。然而,与均相溶液不同,受体分子在细胞膜上的分布是非连续的、动态的,因此细胞表面的受体配体相互作用通常呈现复杂的非线性结合模式。框架核酸作为一类具有确定几何形状的DNA纳米支架,可用于多价配体的偶联,为深入揭示受体配体相互作用机制提供了可靠的工具。利用框架核酸纳米分辨率的可寻址特性,可实现对配体数目、间距及空间构象等参数的精确调控,进而研究细胞表面受体配体的结合特性及影响因素,优化结合条件最终实现高效的分子识别及靶向治疗。本文综述了基于框架核酸的细胞表面受体配体相互作用研究进展,通过探讨细胞表面受体配体相互作用的重要影响因素及生物学应用,对该研究领域的发展前景和未来趋势予以展望。
中图分类号:
引用本文
郭琳洁, 彭红珍, 李江, 王丽华, 诸颖. 基于框架核酸的细胞表面受体配体相互作用研究进展[J]. 应用化学, 2022, 39(10): 1475-1487.
Lin-Jie GUO, Hong-Zhen PENG, Jiang LI, Li-Hua WANG, Ying ZHU. Advances in Receptor‑ligand Interactions on Cell Surface Based on Framework Nucleic Acids[J]. Chinese Journal of Applied Chemistry, 2022, 39(10): 1475-1487.

图1 DNA与配体的非共价连接与共价连接(a) 可卡因酯酶CocE通过生物素亲和素非共价相互作用连接在DNA折纸的指定位点[19]; (b) 带His标签的EGFP蛋白在Ni2+离子存在下,与氮三乙酸(NTA)标记的DNA非共价连接[22]; (c) SPDP、SMCC、SNAP-tag和Halo-tag介导的DNA与配体的共价连接作用机制[16]
Fig.1 Non-covalent and covalent conjugation between DNA and ligands(a) The cocaine-esterase CocE non-covalently linked to the specified site of DNA origami through streptavidin and biotin interactions[19]; (b) EGFP protein with Histag non-covalently linked to NTA-DNA in the presence of Ni2+[22]; (c) SPDP, SMCC, Snap-tag and Halo-tag mediated covalent conjugation[16]
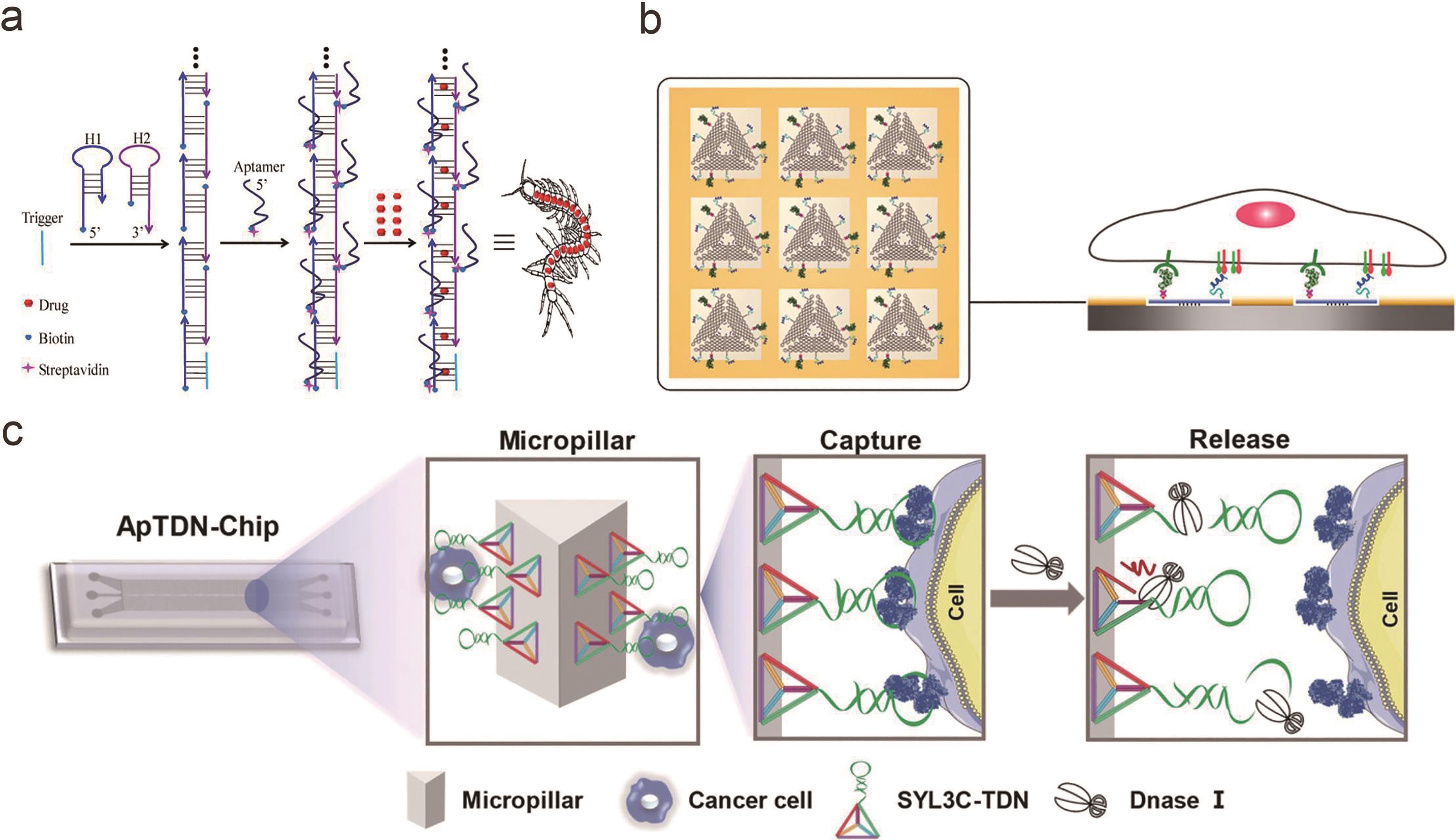
图2 基于框架核酸的受体配体多价相互作用(a) 基于自组装DNA“纳米蜈蚣”的多价核酸适配体[20]; (b) 三角形DNA折纸上修饰多价EGF和A20FMDV2多肽用于癌细胞扩散中受体配体相互作用研究[42]; (c) DNA四面体作为支架在微流体芯片界面上形成多价核酸适配体,对循环肿瘤细胞进行捕获与释放[49]
Fig.2 Multivalent interactions between receptor and ligand based on framework nucleic acids(a) Multivalent aptamers conjugated to self-assembled DNA “nanocentipede”[20]; (b) EGF and A20FMDV2 peptides modified on triangular DNA origami for the study of receptor ligand interactions in cancer cell proliferation[42]; (c) DNA tetrahedron with multivalent aptamers on the microfluidic chip interface for circulating tumor cells capture and release[49]

图3 基于框架核酸的受体配体距离依赖的相互作用(a) DNA三角折纸上不同距离人工抗原表位设计及高速原子力显微镜下抗原抗体相互作用的单分子成像[61]; (b) eOD-GT8抗原以不同数目和间距连接在二十面体及六螺旋DNA折纸表面[62]; (c) 方块折纸两个垂直方向上不同距离caspase-9单体的原子力显微镜成像[63]
Fig.3 Distance-dependent interactions between receptor and ligand based on framework nucleic acids(a) Design of artificial antigen epitopes with different distances on DNA triangular origami and single-molecule imaging of antigen-antibody interactions under high-speed atomic force microscopy[61]; (b) eOD-GT8 antigens attached to icosahedral and six-helix DNA origami surfaces in varying numbers and intervals[62]; (c) Atomic force microscopy imaging of caspase-9 monomer at different distances in two vertical directions on square DNA origami[63]
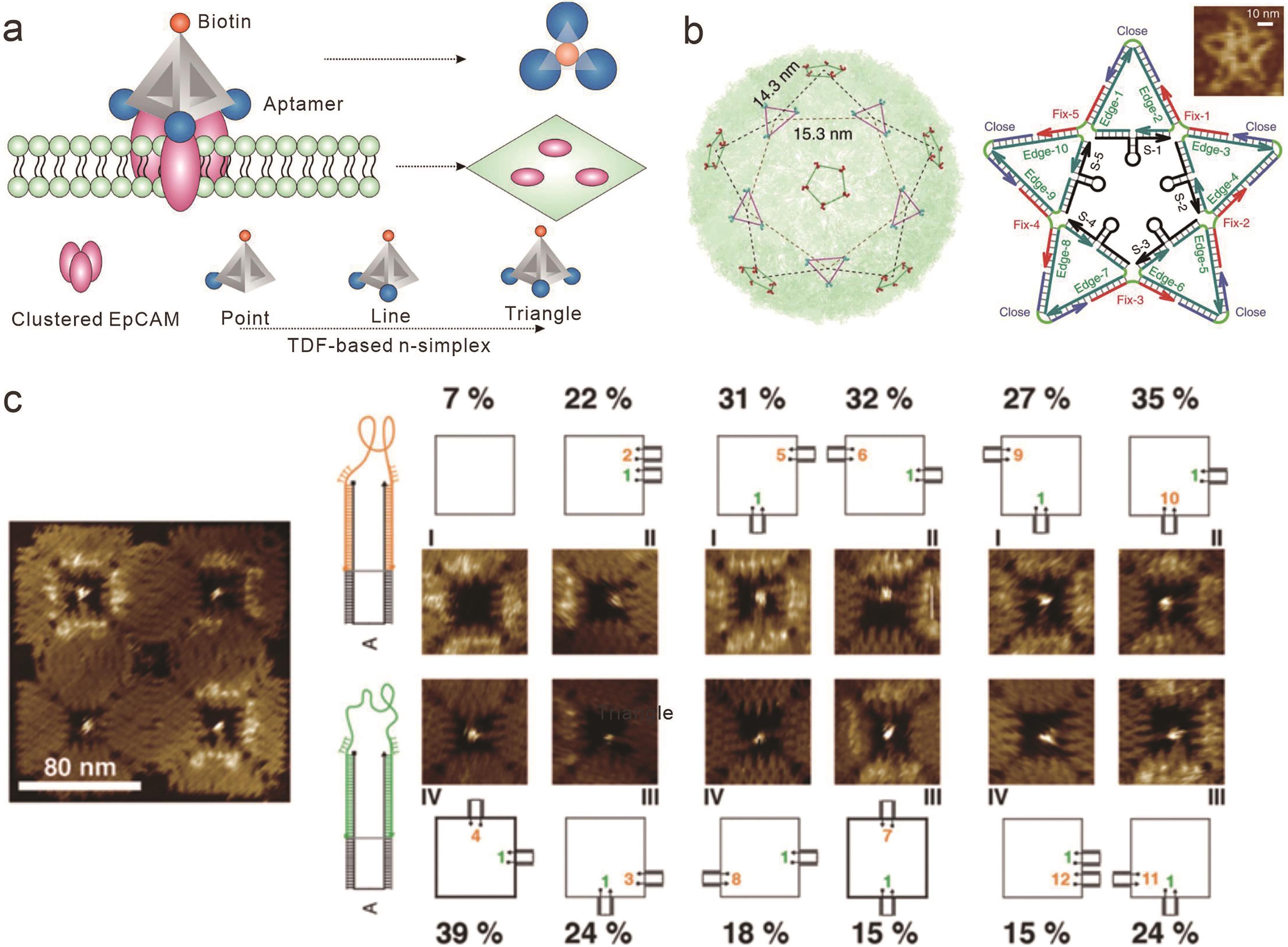
图4 基于框架核酸的受体配体拓扑构象依赖的相互作用(a) 基于DNA四面体的核酸适配体拓扑重排及其与细胞膜上受体结合[67]; (b) 根据登革热病毒表面3价与5价包膜蛋白域Ⅲ(ED3)设计的携带10个ED3核酸适配体的星形DNA结构[69]; (c) 二维DNA折纸的纳米空腔上11种核酸适配体构象变化及其与配体的结合率比较[70]
Fig.4 Topologically conformation dependent interactions between receptor and ligand based on framework nucleic acids(a) Topological rearrangement of aptamers based on DNA tetrahedrons and their binding to cell membrane receptors[67]; (b) Star-shaped DNA structure carrying 10 ED3 aptamers based on surface trivalent and pentavalent envelope protein domain Ⅲ (ED3) of dengue virus[69]; (c) Comparison of conformation changes of 11 aptamers and their binding rates with ligands in nanocavities of two-dimensional DNA origami[70]
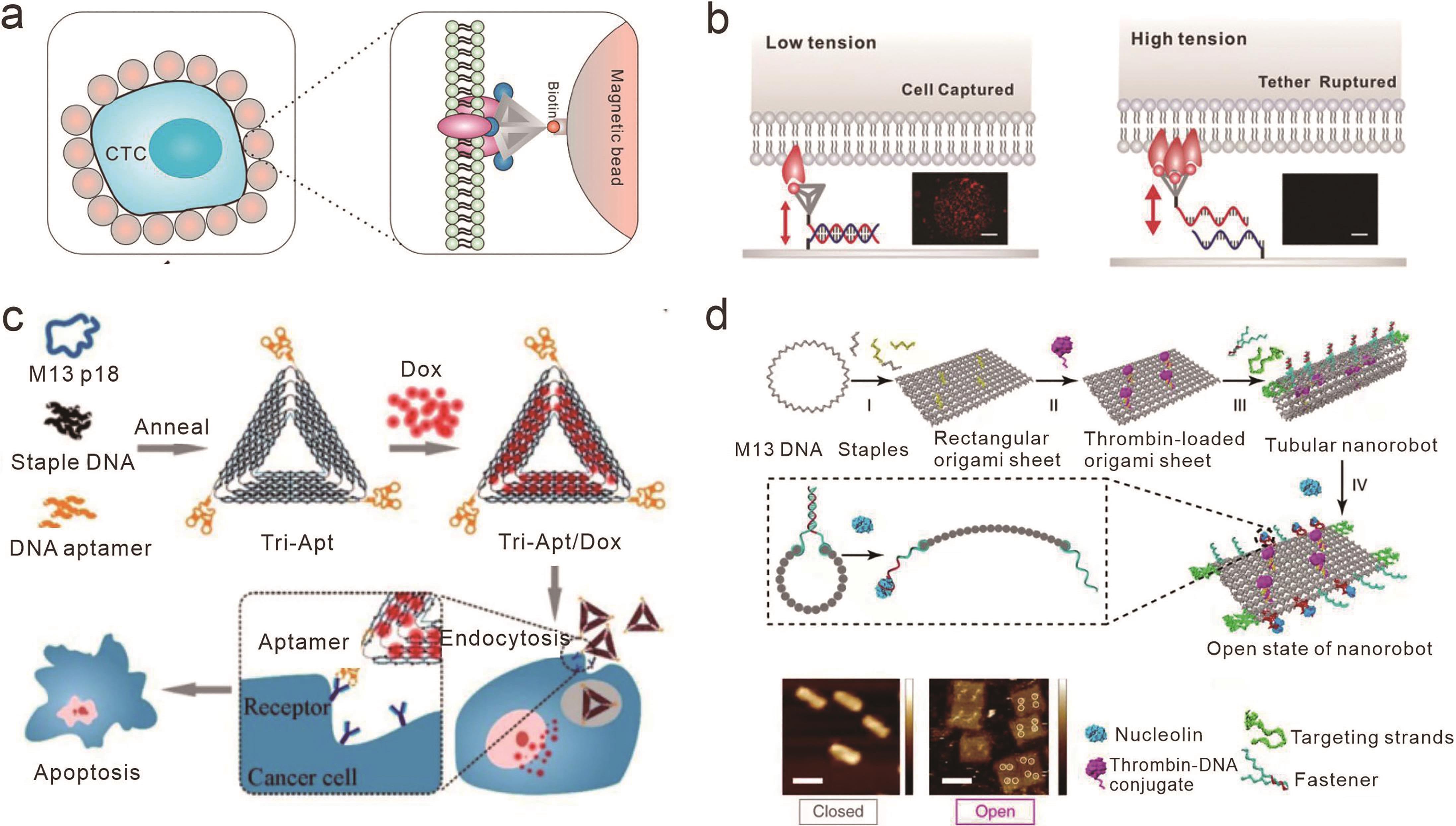
图5 基于框架核酸的受体配体相互作用的生物学应用(a) 基于DNA四面体的多价核酸适配体用于循环肿瘤细胞捕获[67]; (b) 测量拓扑工程DNA四面体编码的多价核酸适配体与细胞表面受体结合力[68]; (c) 多价核酸适配体结合的DNA折纸作为抗癌药物阿霉素(Dox)的靶向递送载体[75]; (d) 连接核酸适配体的DNA纳米机器人用于凝血酶的靶向递送和响应性释放[78]
Fig.5 Biological applications of receptor-ligand interactions based on framework nucleic acids(a) Multivalent aptamers based on DNA tetrahedrons for circulating tumor cell capture[67]; (b) Measuring the binding force between multivalent aptamers and cell surface receptors encoded by topological engineering DNA tetrahedrons[68]; (c) Multivalent aptamer-conjugated DNA origami as a targeted delivery vector for the anticancer drug Doxorubicin[75]; (d) Targeted delivery and responsive release of thrombin by DNA nanorobots linked with aptamers[78]
| 1 | KIESSLING L L, GESTWICKI J E.STRONG L E. Synthetic multivalent ligands in the exploration of cell-surface interactions[J]. Curr Opin Chem Biol, 2000, 4: 696-703. |
| 2 | UEKI R, ATSUTA S, UEKI A, et al. Nongenetic reprogramming of the ligand specificity of growth factor receptors by bispecific DNA aptamers[J]. J Am Chem Soc, 2017, 139(19): 6554-6557. |
| 3 | HUPPA J B, AXMANN M, MORTELMAIER M A, et al. Tcr-peptide-MHC interactions in situ show accelerated kinetics and increased affinity[J]. Nature, 2010, 463(7283): 963-967. |
| 4 | TONG G J, HSIAO S C, CARRICO Z M, et al. Viral capsid DNA aptamer conjugates as multivalent cell-targeting vehicles[J]. J Am Chem Soc, 2009, 131: 11174-11178. |
| 5 | YUAN Q, WU Y, WANG J, et al. Targeted bioimaging and photodynamic therapy nanoplatform using an aptamer-guided g-quadruplex DNA carrier and near-infrared light[J]. Angew Chem Int Ed, 2013, 52(52): 13965-13969. |
| 6 | WU Y, SEFAH K, LIU H, et al. DNA aptamer-micelle as an efficient detection/delivery vehicle toward cancer cells[J]. Proc Natl Acad Sci U S A, 2010, 107(1): 5-10. |
| 7 | SHENG W, CHEN T, TAN W, et al. Multivalent DNA nanospheres for enhanced capture of cancer cells in microfluidic devices[J]. ACS Nano, 2013, 7: 7067-7076. |
| 8 | HUANG Y, CHANG H, TAN W. Cancer cell targeting using multiple aptamers conjugated on nanorods[J]. Anal Chem, 2008, 80: 567-572. |
| 9 | GENG Z, WANG L, LIU K, et al. Enhancing anti-PD-1 immunotherapy by nanomicelles self-assembled from multivalent aptamer drug conjugates[J]. Angew Chem Int Ed, 2021, 60(28): 15459-15465. |
| 10 | YANG Y, SUN X, XU J, et al. Circular bispecific aptamer-mediated artificial intercellular recognition for targeted T cell immunotherapy[J]. ACS Nano, 2020, 14(8): 9562-9571. |
| 11 | GE Z, GU H, LI Q, et al. Concept and development of framework nucleic acids[J]. J Am Chem Soc, 2018, 140(51): 17808-17819. |
| 12 | SEEMAN N C, SLEIMAN H F. DNA nanotechnology[J]. Nat Rev Mater, 2017, 3: 17068. |
| 13 | LIU N, LIEDL T. DNA-assembled advanced plasmonic architectures[J]. Chem Rev, 2018, 118(6): 3032-3053. |
| 14 | OUYANG X, WANG M, GUO L, et al. DNA nanoribbon-templated self-assembly of ultrasmall fluorescent copper nanoclusters with enhanced luminescence[J]. Angew Chem Int Ed, 2020, 59(29): 11836-11844. |
| 15 | ZHANG Q, JIANG Q, LI N, et al. DNA origami as an in vivo drug delivery vehicle for cancer therapy[J]. ACS Nano, 2014, 8: 6633-6643. |
| 16 | KONG G, XIONG M, LIU L, et al. DNA origami-based protein networks: from basic construction to emerging applications[J]. Chem Soc Rev, 2021, 50(3): 1846-1873. |
| 17 | KUZUYA A, KIMURA M, NUMAJIRI K, et al. Precisely programmed and robust 2D streptavidin nanoarrays by using periodical nanometer-scale wells embedded in DNA origami assembly[J]. ChemBioChem, 2009, 10(11): 1811-1815. |
| 18 | NUMAJIRI K, KIMURA M, KUZUYA A, et al. Stepwise and reversible nanopatterning of proteins on a DNA origami scaffold[J]. Chem Commun, 2010, 46(28): 5127-5129. |
| 19 | MALLIK L, DHAKAL S, NICHOLS J, et al. Electron microscopic visualization of protein assemblies on flattened DNA origami[J]. ACS Nano, 2015, 9: 7133-7141. |
| 20 | LI W, YANG X, HE L, et al. Self-assembled DNA nanocentipede as multivalent drug carrier for targeted delivery[J]. ACS Appl Mater Interfaces, 2016, 8(39): 25733-25740. |
| 21 | OUYANG X, DE STEFANO M, KRISSANAPRASIT A, et al. Docking of antibodies into the cavities of DNA origami structures[J]. Angew Chem Int Ed, 2017, 56(46): 14423-14427. |
| 22 | SHEN W, ZHONG H, NEFF D, et al. NTA directed protein nanopatterning on DNA origami nanoconstructs[J]. J Am Chem Soc, 2009, 131: 6660-6661. |
| 23 | YAMAZAKI T, HEDDLE J G, KUZUYA A, et al. Orthogonal enzyme arrays on a DNA origami scaffold bearing size-tunable wells[J]. Nanoscale, 2014, 6(15): 9122-9126. |
| 24 | 黄子珂, 刘超, 付强强, 等. 核酸适配体荧光探针在生化分析和生物成像中的研究进展[J]. 应用化学, 2018, 35(1): 28-39. |
| HUANG Z K, LIU C, FU Q Q, et al. Aptamer-based fluorescence probe for bioanalysis and bioimaging[J]. Chinese J Appl Chem, 2018, 35(1): 28-39. | |
| 25 | CHHABRA R, SHARMA J, KE Y, et al. Spatially addressable multiprotein nanoarrays templated by aptamer-tagged DNA nanoarchitectures[J]. J Am Chem Soc, 2007(129): 10304-10305. |
| 26 | FU J, LIU M, LIU Y, et al. Interenzyme substrate diffusion for an enzyme cascade organized on spatially addressable DNA nanostructures[J]. J Am Chem Soc, 2012, 134(12): 5516-5519. |
| 27 | FISHER P D E, SHEN Q, AKPINAR B, et al. A programmable DNA origami platform for organizing intrinsically disordered nucleoporins within nanopore confinement[J]. ACS Nano, 2018, 12(2): 1508-1518. |
| 28 | NAKATA E, DINH H, NGO T A, et al. A modular zinc finger adaptor accelerates the covalent linkage of proteins at specific locations on DNA nanoscaffolds[J]. Chem Commun, 2015, 51(6): 1016-1019. |
| 29 | KOSSMANN K J, ZIEGLER C, ANGELIN A, et al. A rationally designed connector for assembly of protein-functionalized DNA nanostructures[J]. ChemBioChem, 2016, 17(12): 1102-1106. |
| 30 | SACCA B, MEYER R, ERKELENZ M, et al. Orthogonal protein decoration of DNA origami[J]. Angew Chem Int Ed, 2010, 49(49): 9378-9383. |
| 31 | ILIUK A B, HU L, TAO W A. Aptamer in bioanalytical applications[J]. Anal Chem, 2011, 83(12): 4440-4452. |
| 32 | TAN W, DONOVAN M J, JIANG J, Aptamers from cell-based selection for bioanalytical applications[J]. Chem Rev, 2013, 113(4): 2842-2862. |
| 33 | MAMMEN M, CHOI S, WHITESIDES G M. Polyvalent interactions in biological systems: implications for design and use of multivalent ligands and inhibitors[J]. Angew Chem Int Ed, 1998, 37: 2754-2794. |
| 34 | FASTING C, SCHALLEY C A, WEBER M, et al. Multivalency as a chemical organization and action principle[J]. Angew Chem Int Ed, 2012, 51(42): 10472-10498. |
| 35 | CZAJKOWSKYA D M, SHAO Z. The human IgM pentamer is a mushroom-shaped molecue with a flexural bias[J]. PNAS, 2009, 106: 14960-14965. |
| 36 | HERNANDEZ-LOPEZ R A, YU W, CABRAL K A, et al. T cell circuits that sense antigen density with an ultrasensitive threshold[J]. Science, 2021, 371: 1166-1171. |
| 37 | LIN M, ZHANG J, WAN H, et al. Rationally designed multivalent aptamers targeting cell surface for biomedical applications[J]. ACS Appl Mater Interfaces, 2021, 13(8): 9369-9389. |
| 38 | ZHU G, HU R, ZHAO Z, et al. Noncanonical self-assembly of multifunctional DNA nanoflowers for biomedical applications[J]. J Am Chem Soc, 2013, 135(44): 16438-16445. |
| 39 | LIU X, YAN H, LIU Y, et al. Targeted cell-cell interactions by DNA nanoscaffold-templated multivalent bispecific aptamers[J]. Small, 2011, 7(12): 1673-1682. |
| 40 | ROTHEMUND P W. Folding DNA to create nanoscale shapes and patterns[J]. Nature, 2006, 440(7082): 297-302. |
| 41 | 付衍明, 张钊, 李璨, 等. DNA折纸术研究进展[J]. 应用化学, 2010, 27 (2): 125-131. |
| FU Y M, ZHANG Z, LI C, et al. Recent progress in DNA origami[J]. Chinese J Appl Chem, 2010, 27(2): 125-131. | |
| 42 | HUANG D, PATEL K, PEREZ-GARRIDO S, et al. DNA origami nanoarrays for multivalent investigations of cancer cell spreading with nanoscale spatial resolution and single-molecule control[J]. ACS Nano, 2019, 13(1): 728-736. |
| 43 | HAWKES W, HUANG D, REYNOLDS P, et al. Probing the nanoscale organisation and multivalency of cell surface receptors: DNA origami nanoarrays for cellular studies with single-molecule control[J]. Faraday Discuss, 2019, 219(0): 203-219. |
| 44 | DUTTA P K, ZHANG Y, BLANCHARD A T, et al. Programmable multivalent DNA-origami tension probes for reporting cellular traction forces[J]. Nano Lett, 2018, 18(8): 4803-4811. |
| 45 | GOODMAN R P, BERRY R M, TURBERFIELD A J. The single-step synthesis of a DNA tetrahedron[J]. Chem Commun, 2004(12): 1372-1373. |
| 46 | GOODMAN R P, SCHAAP A T, TARDIN C F, et al. Rapid chiral assembly of rigid DNA building blocks for molecular nanofabrication[J]. 2005, 310: 1661-1665. |
| 47 | ZHANG T, CUI W, TIAN T, et al. Progress in biomedical applications of tetrahedral framework nucleic acid-based functional systems[J]. ACS Appl Mater Interfaces, 2020, 12(42): 47115-47126. |
| 48 | LI J, PEI H, ZHU B, et al. Self-assembled multivalent DNA nanostructures for noninvasive intracellular delivery of immunostimulatory CpG oligonucleotides[J]. ACS Nano, 2011, 5: 8783-8789. |
| 49 | ZHANG J, LIN B, WU L, et al. DNA nanolithography enables a highly ordered recognition interface in a microfluidic chip for the efficient capture and release of circulating tumor cells[J]. Angew Chem Int Ed, 2020, 59(33): 14115-14119. |
| 50 | LI J, DAI J, JIANG S, et al. Encoding quantized fluorescence states with fractal DNA frameworks[J]. Nat Commun, 2020, 11(1): 2185 |
| 51 | XIE M, GUO L, XING S, et al. Cell imaging with multi-color DNA framework probes[J]. Chem Commun, 2021, 57(86): 11318-11321. |
| 52 | SIMONS K, TOOMRE D. Lipid rafts and signal transduction[J]. Nat Rev, 2000, 1: 31-41. |
| 53 | GOMES DE CASTRO M A, WILDHAGEN H, SOGRATE-IDRISSI S, et al. Differential organization of tonic and chronic B cell antigen receptors in the plasma membrane[J]. Nat Commun, 2019, 10(1) :820. |
| 54 | DEY S, FAN C, GOTHELF K V, et al. DNA origami[J]. Nat Rev Methods Primers, 2021, 1(1): 13. |
| 55 | RINKER S, KE Y, LIU Y, et al. Self-assembled DNA nanostructures for distance-dependent multivalent ligand-protein binding[J]. Nat Nanotechnol, 2008, 3(7): 418-422. |
| 56 | SHAW A, LUNDIN V, PETROVA E, et al. Spatial control of membrane receptor function using ligand nanocalipers[J]. Nat Methods, 2014, 11(8): 841-846. |
| 57 | DIEBOLDER C A, BEURSKENS F J, JONG R N, et al. Complement is activated by IgG hexamers assembled at the cell surface[J]. Science, 2014, 343: 1260-1264. |
| 58 | SONDERMANN P, HUBER R, OOSTHUIZEN V, et al. The 3.2-Å crystal structure of the human IgG fc fragment-fcγriii complex[J]. Nature, 2000, 406: 267-273. |
| 59 | QIAO S, KOBAYASHI K, JOHANSEN F E, et al. Dependence of antibody-mediated presentation of antigen on FcRn[J]. Proc Nat Acad Sci, 2008, 105: 9337-9342. |
| 60 | SHAW A, SANDLIE I, MICHAELSEN T E, et al. Binding to nanopatterned antigens is dominated by the spatial tolerance of antibodies[J]. Nat Nanotechnol, 2019, 14: 184-190. |
| 61 | ZHANG P, LIU X, LIU P, et al. Capturing transient antibody conformations with DNA origami epitopes[J]. Nat Commun, 2020, 11(1): 3114. |
| 62 | VENEZIANO R, MOYER T J, STONE M B, et al. Role of nanoscale antigen organization on B-cell activation probed using DNA origami[J]. Nat Nanotechnol, 2020, 15(8): 716-723. |
| 63 | ROSIER B, MARKVOORT A J, GUMI AUDENIS B, et al. Proximity-induced caspase-9 activation on a DNA origami-based synthetic apoptosome[J]. Nat Catal, 2020, 3(3): 295-306. |
| 64 | ZHANG K, DENG R, SUN Y, et al. Reversible control of cell membrane receptor function using DNA nano-spring multivalent ligands[J]. Chem Sci, 2017, 8(10): 7098-7105. |
| 65 | HARRISON S C. Looking inside adenovirus[J]. Science, 2010, 329: 1026-1027. |
| 66 | CAI H, MULLER J, DEPOIL D, et al. Full control of ligand positioning reveals spatial thresholds for T cell receptor triggering[J]. Nat Nanotechnol, 2018, 13: 610-617. |
| 67 | LI M, DING H, LIN M, et al. DNA framework-programmed cell capture via topology-engineered receptor-ligand interactions[J]. J Am Chem Soc, 2019, 141(47): 18910-18915. |
| 68 | YIN F, LI M, MAO X, et al. DNA framework-based topological cell sorters[J]. Angew Chem Int Ed, 2020, 59(26): 10406-10410. |
| 69 | KWON P S, REN S, KWON S J, et al. Designer DNA architecture offers precise and multivalent spatial pattern-recognition for viral sensing and inhibition[J]. Nat Chem, 2020, 12(1): 26-35. |
| 70 | RAFAT A A, SAGREDO S, THALHAMMER M, et al. Barcoded DNA origami structures for multiplexed optimization and enrichment of DNA-based protein-binding cavities[J]. Nat Chem, 2020, 12: 852-859. |
| 71 | ZHONG L, CAI S, HUANG Y, et al. DNA octahedron-based fluorescence nanoprobe for dual tumor-related mrnas detection and imaging[J]. Anal Chem, 2018, 90(20): 12059-12066. |
| 72 | XUE C, ZHANG S, LI C, et al. Y-shaped backbone-rigidified triangular DNA scaffold-directed stepwise movement of a dnazyme walker for sensitive microrna imaging within living cells[J]. Anal Chem, 2019, 91(24): 15678-15685. |
| 73 | LI Q, ZHAO D, SHAO X, et al. Aptamer-modified tetrahedral DNA nanostructure for tumor-targeted drug delivery[J]. ACS Appl Mater Interfaces, 2017, 9(42): 36695-36701. |
| 74 | SHI S, FU W, LIN S, et al. Targeted and effective glioblastoma therapy via aptamer-modified tetrahedral framework nucleic acid-paclitaxel nanoconjugates that can pass the blood brain barrier[J]. Nanomedicine, 2019, 21: 102061. |
| 75 | CAO M, SUN Y, XIAO M, et al. Multivalent aptamer-modified DNA origami as drug delivery system for targeted cancer therapy[J]. Chem Res Chinese Univ, 2019, 36(2): 254-260. |
| 76 | GE Z, GUO L, WU G, et al. DNA origami-enabled engineering of ligand-drug conjugates for targeted drug delivery[J]. Small, 2020, 16(16): e1904857. |
| 77 | ZHANG P, FISCHER A, OUYANG Y, et al. Aptamer-modified DNA tetrahedra-gated metal-organic framework nanoparticle carriers for enhanced chemotherapy or photodynamic therapy[J]. Chem Sci, 2021, 12(43): 14473-14483. |
| 78 | LI S, JIANG Q, LIU S, et al. A DNA nanorobot functions as a cancer therapeutic in response to a molecular trigger in vivo[J]. Nat Biotech, 2018, 36: 258-264. |
| [1] | 闫美玲, 彭红珍, 左婷婷, 田甜, 诸颖, 孙艳红. 四面体框架核酸对脑靶向肽分子的可控组装及性能[J]. 应用化学, 2022, 39(10): 1501-1509. |
| [2] | 韦国兵,孔德荣,徐慧慧,殷赵江,张晶,崔汉峰,洪年,杨婕,熊魏,刘文明,郭倩文,程林,樊浩. 基于主客体识别作用构建的电化学发光传感器检测丹参中汞离子[J]. 应用化学, 2019, 36(5): 595-602. |
| [3] | 黄秋颖, 林肖漪, 王勇, 朱永朝. 波浪形一维链状镉配合物的合成、结构及分子识别性能[J]. 应用化学, 2017, 34(9): 1093-1098. |
| [4] | 朱江, 苟文秀, 王孝敏, 陈立东, 冷茹冰, 姜春杰. 萘磺酸型离子配合物的合成及其分子识别性能[J]. 应用化学, 2017, 34(1): 105-110. |
| [5] | 张铁莉, 柴国平, 赫彩霞, 王磊. 对羟基苯甲酸印迹聚合物微粒的制备及其固相萃取应用[J]. 应用化学, 2016, 33(2): 229-237. |
| [6] | 王素素, 张月, 李辉, 许苗苗. 芦丁-槲皮素双模板印迹聚合物的制备、表征及识别[J]. 应用化学, 2015, 32(11): 1290-1298. |
| [7] | 曹林交, 高保娇, 胡伟民. 生物碱分子表面印迹聚合物材料的设计与制备及其分子识别特性[J]. 应用化学, 2014, 31(12): 1390-1398. |
| [8] | 侯长军, 张玉婵, 黄舜, 刘振, 沈才洪, 张宿义, 霍丹群. 卟啉及其衍生物对生物分子的识别研究进展[J]. 应用化学, 2011, 28(09): 977-985. |
| [9] | 吴粦华, 拓宏桂, 钟春龙. 2-羟基-1-萘甲醛缩-2-萘甲酰腙对Zn2+的选择性识别[J]. 应用化学, 2010, 27(06): 632-636. |
| [10] | 许志锋,吴双,吴小辉,邝代治,张复兴,王剑秋. 二茂铁甲酸分子印迹聚合物的制备和识别机理、结合特性[J]. 应用化学, 2010, 27(04): 384-389. |
| [11] | 曹玺珉, 廖玲, 杜黎明. 依诺沙星分子印迹聚合物的分子识别[J]. 应用化学, 2008, 25(1): 43-47. |
| [12] | 李来生, 葛小辉, 黄志兵, 李艳平. 光谱法研究羟基葫芦[6]脲与对氨基苯磺酸的分子识别作用[J]. 应用化学, 2006, 23(7): 747-752. |
| [13] | 邴乃慈, 许振良, 杨座国, 王学军, 冯建立. 热聚合制备左氧氟沙星分子印迹聚合物的分子识别特性[J]. 应用化学, 2006, 23(10): 1085-1089. |
| [14] | 刘慧君, 项伟中, 徐伟箭, 熊远钦, 卢彦兵. 磺胺二甲嘧啶分子印迹聚合物的合成及其识别性能[J]. 应用化学, 2005, 22(1): 83-86. |
| [15] | 尹伟, 张秀莲, 张迈生. 氮杂对环吩CP66与萘衍生物超分子体系的研究[J]. 应用化学, 2001, 18(6): 457-461. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||